| 技術名稱 | 適用於次世代定序識別基因變體之系統晶片 | ||
|---|---|---|---|
| 計畫單位 | 國立陽明交通大學 | ||
| 計畫主持人 | 洪瑞鴻 | ||
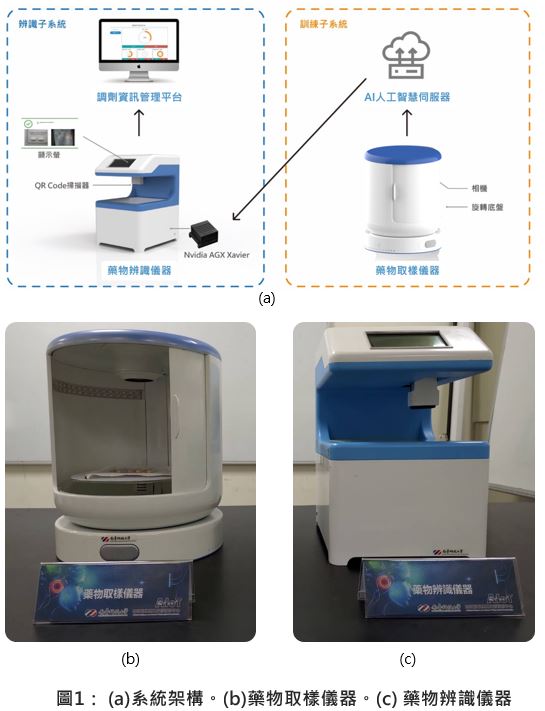
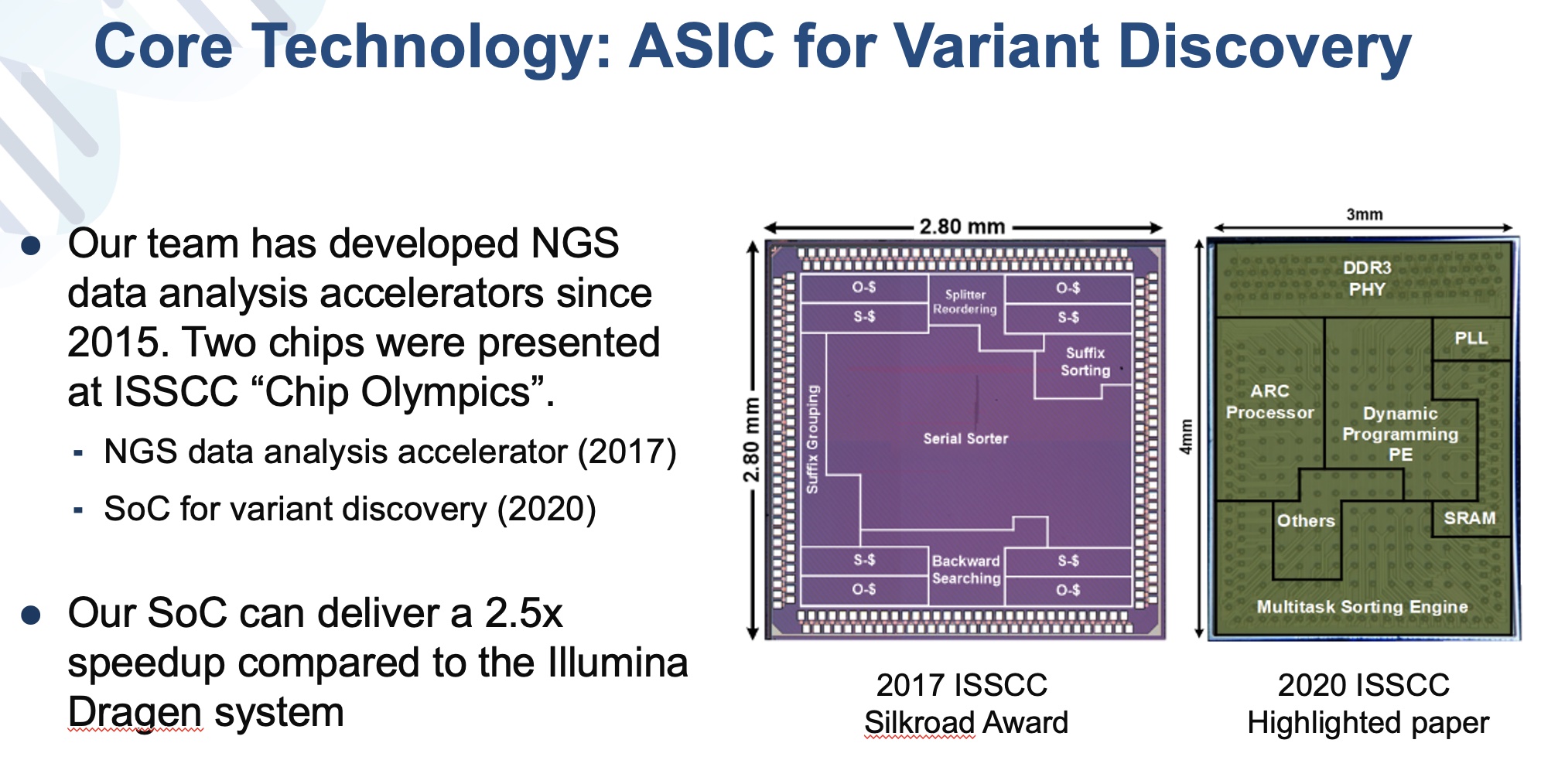
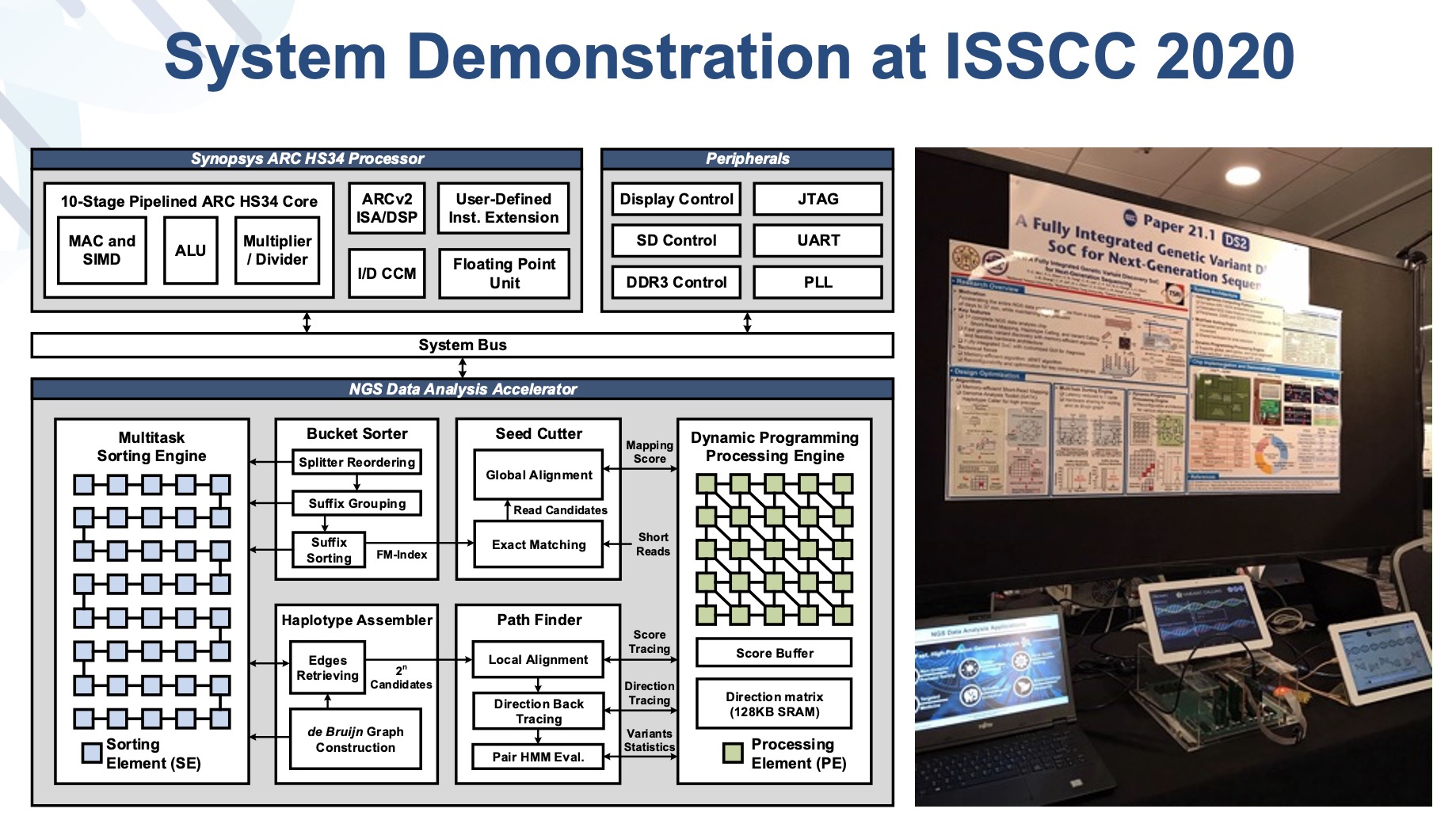
| 技術簡介 | 本研究提出一個新式整合次世代基因定序資料分析之變體識別系統單晶片架構,同時此平台為完全可攜式,便於攜帶至醫療比對,並透過GATK演算法進行序列重組以建立半倍體。最後比對半倍體與參考基因序列並記錄變異資訊。場所與研究地點以進行即時診斷。 |
||
| 技術影片 | |||
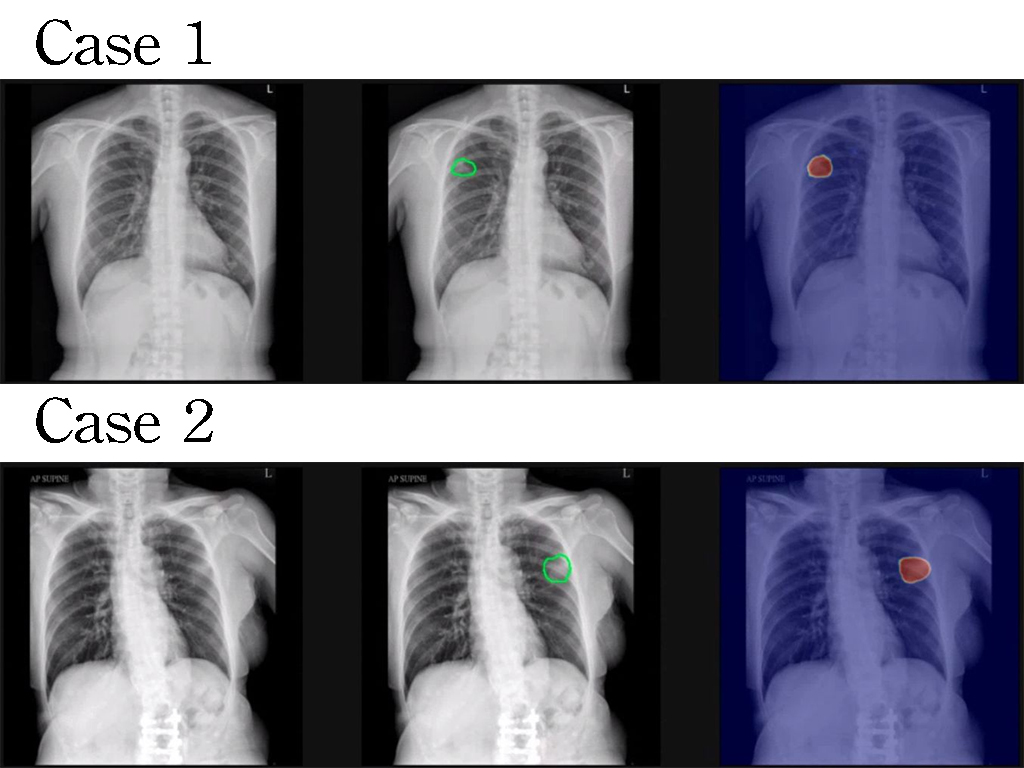
| 科學突破性 | 現有次世代基因定序資料分析電路多研究部分步驟優化。本作品整合完整資料分析流程,達到全世界唯一完整且最高速之資料分析系統,相較於過往之高階GPU平台,此作品達到66倍的加速、並在能量效率與面積效率上亦達到超過15,000與3,000的增益。此設計透過美國FDA所提供之測試資料,達到99.6的準確度。 |
||
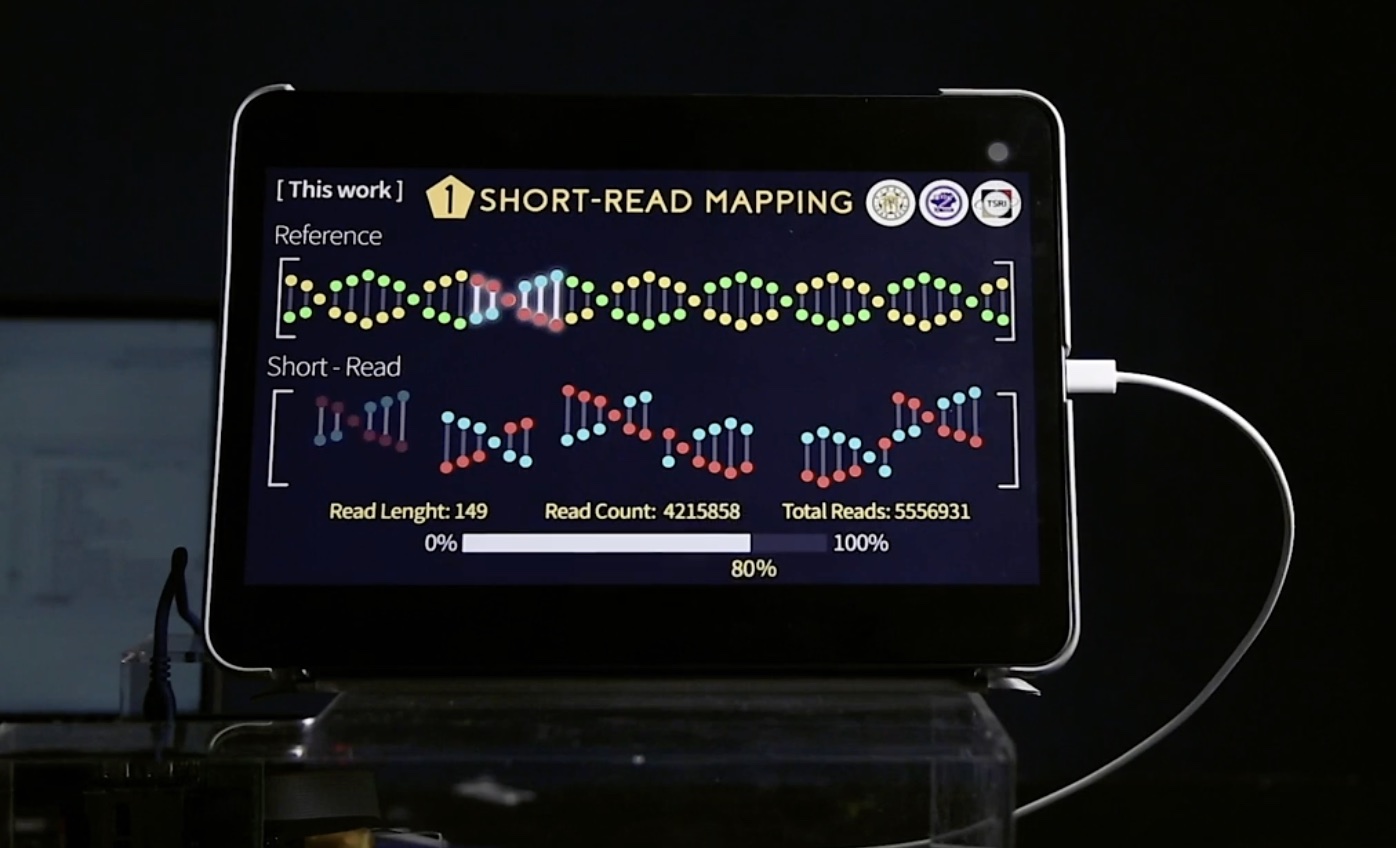
| 產業應用性 | 次世代基因定序市場之未來產值被產業界公認為將會指數上升。然而,定序後續的資料分析過於耗時,需花費超過數天至一個禮拜以上。本團隊致力於開發適用於次世代定序識別基因變體之系統單晶片,大幅加速基因定序資料分析運算,將運算時間縮短至四十分鐘內,且為可攜式以便於攜帶至醫療場所與研究地點以利醫生進行即時診斷。 |
||
| 媒合需求 | 天使投資人、策略合作夥伴 |
||
| 關鍵字 | 次世代基因定序資料分析 高速硬體排序 超大型積體電路與系統 系統晶片 | ||
- 聯絡人
- 洪瑞鴻
- 電子信箱
- jhh@cs.nctu.edu.tw